Correspondence: Tomi Heinonen,
Ragnar Granit Institute, Tampere University of Technology,
P.O. Box 692, FIN-33101 Tampere, Finland. E-mail: Tomi.Heinonen@Nokia.com,
phone +358 50 3738 611, fax +358 3 247 4013
1. Introduction
Most of the new medical imaging devices are digital enabling
computerized analysis of the produced images. In general,
computerized image analysis corresponds for image enhancement,
image restoration, image filtering, and image measurement.
However, section images produced by e.g., Magnetic Resonance
Imaging (MRI) and Computed Tomography (CT) enable a variety
of new image analysis applications. The most important from
these is segmentation [Heinonen, 1999].
Segmentation corresponds to a decomposition of a scene
into its components. On medical images, the segmented structures
can be e.g., different organs and tissues. The main applications
(see Fig. 1) of segmented images are volumetry (i.e., volume
analysis of segmented structures), computer models (e.g.,
brain models and thorax models), and three-dimensional visualization
(e.g., separate displaying of segmented structures). Some
examples of these main applications are presented in the
Fig. 2.

Figure 1. Segmentation
is a central technique in enabling volumetric analysis of
medical images, construction of computer models for thorax
and brain, and 3D visualization. Together with biosignals
and computer models, it is possible to calculate electrical
fields inside the body and present the results as multimodal
visualization.



Figure 2. On the left
side, several MRI slices have been segmented in order to
carry out volumetry of Multiple Sclerosis plaques and fluid
spaces. On middle, one segmented slice of a thorax model
is presented together with simulation results. On the rigth
side, various 3D visualization modes of segmented images.
2. Segmentation
In general, segmentation involves several stages, which
are Image Enhancement, Feature Extraction, Segmentation,
and Classification (see Fig. 4). All the stages are not
always necessary and sometimes segmentation can lead directly
to classification and vice versa. Image enhancement is capable
of improving the image appearance (e.g., removal of noise,
contrast stretching, etc), feature extraction emphasizes
regions of interest (e.g., emphasizing borders), segmentation
separates emphasized structures from background, and classification
recognizes/classifies them [Jain A, 1989].

Figure 3. Segmentation
stages; Image enhancement is capable of improving the image
appearance (e.g., removal of noise, contrast stretching,
etc), feature extraction emphasizes regions of interest
(e.g., emphasizing berders), segmentation separates emphasized
structures from background, and classification recognizes/classifies
them.
Segmentation techniques can be classified using several
criteria: Techniques detecting borders of organs/tissues
are Boundary based techniques, techniques detecting
similar intensities or textures are Region based
techniques, Hybrid techniques utilize different methods
(e.g., Boundary + Region) simultaneously with model parameters
or multispectral images, and Classification applies
common pattern recognition methodology to recognize structures.

Figure 4. Various
Segmentation techniques – segmentation can lead to
classification and vice versa.
Different segmentation techniques produce different results:
Boundary based segmentation provides models for edges and
boundaries, which are often coded by chain links. Region
based segmentation produces groups of voxels or just labels
all the voxels in the image set. If the aim of the segmentation
is to reconstruct a model to be used in simulations, boundaries
are trivial to convert to labeled voxels (as long as the
contours are closed), but converting labeled voxels to contours
is more complex, hence it should keep in the focus what
type of segmentation should be selected in particular applications.
In addition, some techniques are sensitive to noise; therefore
the quality of images also directs the selection of segmentation
techniques.

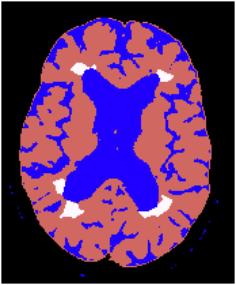
Figure 5. On the left
side; boundary based segmentation result. Scalp and cortex
are surrounded by chain links. On the right side;region
based segmentation result representing the brain, cerebrospinal
fluid and multiple sclerosis plaques.
2.1. Boundary Based Segmentation
The simplest Boundary based technique is manual tracing,
which is still used in several clinical applications. The
aim is to use a pointer (e.g., mouse) and operator’s
visual perception to mark boundaries between structures.
This technique is laborious compared to other techniques
but ensures accurate results. During the last two decades,
this technique has greatly evolved towards edge detection
and automatics. Conventional boundary-based segmentation
techniques utilize different gradient operations [Jain,
1989]. Combining this technique with thresholding, results
in a binary image emphasizing edges. To detect the edges,
several simple techniques such as contour following, edge
linking, heuristic graph search, dynamic programming, and
shortest spanning trees [Kwok et al., 1997] have been developed.
Quite often, conventional methods result in false or broken
edges, due to complex and noisy images. In order to solve
this, techniques called whole boundary methods based on
spatial gradient features near boundaries have been studied
[Bomans et al., 1990; Chakraborty et al., 1996; Yezzi et
al., 1997]. These techniques yield better results. The use
of gradients together with watershed transformation and
morphological operators increase the performance of the
boundary methods [Wang 1997]. Further, statistical techniques
enable relatively effective edge detection regardless of
noise [Thune et al., 1997].
Common feature for most of the boundary methods is the
sensitivity to noise. In addition, complex structures (e.g.,
white matter of the brain) are difficult to surround using
contours due to great number of separate contours.
2.2. Region Based Segmentation
Thresholding is the most commonly used segmentation method.
It is based on homogenous regions instead of contours. It
utilizes amplitude segmentation to find voxel groups of
similar intensity [Sahoo et al., 1988]. Such procedure can
be classified as manual, semi-automatic or automatic depending
on the segmentation application and the definition of threshold
coefficients. Because particular tissues and anatomical
structures usually appear in similar intensities, thresholding
is often applied as a feature extractor. However, when source
images are enhanced or particular multi-spectral images
are applied, it is possible that only the regions of interest
appear in some constant intensity range, hence enabling
automatic segmentation (e.g., segmentation of the bone from
CT image set). Thresholding coefficients are often obtained
from intensity histograms manually or using some algorithms
to find peaks and valleys. The criteria how to choose appropriate
coefficients depends on the segmentation application. One
common criterion is to choose the coefficients so that the
resulting image resembles the original image accurately.
For this purpose, advanced methods have been developed based
on histogram entropy and 2D histograms, which employ spatial
information along with pixel intensities [Pun, 1981]. Such
algorithms have evolved during the last years pursuing better
quality [Sahoo et al., 1997], performance [Gong et al.,
1998], and adaptivity to poor signal to noise ratio [Li
et al., 1997]. Also methods based on measures, other than
histograms, have been developed employing statistical information
for spatial occurrences of pixel intensities [Ramac et al.,
1997].
The method of clustering is based on partitioning an image
to regions of similar features, such as shapes, textures,
and intensities. Clustering has been applied in numerous
projects concerning segmentation of multi-modal medical
images, such as MR images of different pulse sequences [Taxt
et al., 1994]. New clustering techniques involve complex
mathematics and statistical analysis in order to increase
the performance and pattern recognition properties [Yegnanarayana
et al., 1997].
Region growing is maybe the most versatile method in medical
image segmentation. It operates by merging neighboring pixels
of similar features [Jain, 1989]. Usually the ‘seed’
is defined interactively, but in some cases prior knowledge
of seed locations can be applied. The region growing process
can be implemented either in 2D or 3D depending on the source
images. The basic implementation of region growing applies
pixel intensities in the decision of pixel merging; if the
intensity of neighbor pixel is similar to the seed pixel,
the pixels are merged. Image intensities are rarely homogenous
and hence more advanced implementations of region growing
include statistical or geometrical analysis, such as minimum
variance [Revol et al., 1997]. Also conventional methods,
such as ‘split and merge’ techniques and thresholding
have been used as feature extractors to produce images with
uniform intensities [Yong et al., 1986]. Region growing
operation is then applied to these images. Some new region
growing techniques have been developed to operate without
seed parameters [Mehnert et al., 1997], and to include classification.
Region-based techniques are not as sensitive to noise as
boundary based methods, hence enabling effective segmentation
of noisy images. However, boundary based techniques rely
on changes in the gray level rather than their actual values
enabling effective boundary segmentation for regions of
varying intensity. Therefore several research projects are
aiming to combine boundary and region based segmentation
[Chakraborty et al., 1996; Tabb et al., 1997].
2.3. Other Techniques
It is also possible to analyze images statistically applying
feature variables or vectors (e.g., shape, texture, intensity,
similarity, etc). This type of computed parameters are compared
model parameters and thereafter classified. Different image
processing techniques, such as thresholding can be utilized
as a feature extractor. Other new promising techniques are
Fuzzy logic [Caillol et al., 1997], neural networks [Lee
et al., 1997], fractal algorithms [Neil et al., 1997], and
Wavelets.
Also multispectral images and more than one modality at
a time can be utilized. For example, MRI scan of the head
can be automatically segmented from T1 and T2 weighted images,
because bone appears black on both image modalities. In
the case of thorax, the situation is more complicated hence
reliable automatic segmentation is still far away.
3. Discussion
Even though segmentation is useful in medical practice
it has numerous difficulties; human anatomy varies a lot
and pathological lesions increase the complexity and decrease
the predictability of the anatomy, hence automatic segmentation
is not always reliable. On the other hand, manual and semiautomatic
approaches suffer from variability; inter and intra observer
studies have demonstrated great variability, especially
when small structures are regions of interests.
Other problems are associated with artifacts and noise
on the images. It is quite often the case, that patient
images are noisy due to patient movement, because imaging
scans require relatively long time. Most of the segmentation
techniques are sensitive to noise and therefore results
are not accurate – or processing can require hours.
In general, computer-processing capabilities double every
1.5 year. Therefore demanding segmentation algorithms will
be useful in near future, leading to accurate segmentation.
Also new promising technologies associated with Fuzzy logic,
neural networks, and computer vision systems can provide
useful solutions for medical image processing.
The development of MRI devices has lead to several new
imaging sequences capable of emphasizing particular tissues
and conditions (e.g., Flair imaging, which can be applied
in Multiple Sclerosis studies). It is in prospect, that
combination of different imaging sequences can enable accurate
automatic segmentation. It is probably possible in the future
to develop such imaging sequences capable of visualizing
electrical properties of the tissues. In this case, segmentation
would not be required in model constructions.
References
Bomans M, Hohne K, Tiede U, Riemer M. 3-D Segmentation
of MR images of the head for 3-D display. IEEE Transactions
on Medical Imaging,p 9:177-183, 1990.
Caillol H, Pieczynski W, Hillion A. Estimation of fuzzy
gaussian mixture and unsupervised statistical image segmentation.
IEEE Transactions on image processing, 6(3):425-436,
1997.
Chakraborty A, Staib L, Duncan J. Deformable boundary finding
in medical images by gradient and region information. IEEE
Transactions on medical imaging, 15(6): 859-870, 1996.
Gong J, Li L, Chen W. Fast recursive algorithm for two-dimensional
thresholding. Pattern recognition, 31(3):295-300,
1998.
Heinonen T. Application of magnetic resonance image segmentatin
in neurology. PhD Thesis, Tampere University of Technology,
Tampere, 1999.
Jain A. Fundamentals of digital image processing. Prentice-Hall
International, Englewoods Cliffs, USA. 1989.
Kwok S, Constantinides A. A fast recursive shortest spanmning
tree for image segmentation and edge detection. IEEE
Transaction on image processing 6(2); 328-332, 1997.
Lee J, Chen C, Sun Y, Tseng G. Occluded object recognition
using multiscale features and hopfield neural networks.
Pattern recognition 30(1): 113-122, 1997.
Li L, Gong J, Chen W. Grey-level image thresholding based
on Fisher linear projection of two-dimensiona histogram.
Pattern recognition 30(5): 743-749, 1997.
Mehnert A, Jackway P. An improved seeded region growing
algorithm. Pattern recognition letters 18:1065-1071,
1997.
Neil G, Curtis K. Shape recognition using fractal geometry.
Pattern recognition 30(12):1957-1969.
Ramac L, Varshney P. Image thresholding based on Ali-Silvey
distance measures. Pattern recognition 30(7):1161-1174,
1997.
Revol C, Jourlin M. A new minimum variance region growing
algorithm for image segmentation. Pattern recognition
letters 18:249-258, 1997.
Sahoo P, Soltani S, Wong A, Chen Y. A survey of the thresholding
techniques. Computing vision and graphics in image processing
41:233-260, 1988.
Sahoo P, Wilkins C, Yeager J. Threshold selection using
Renyi’s entropy. Pattern recognition 30(1):71-84,
1997.
Tabb M, Ahuja N. Multiscale image segmentation by integrated
edge and region detection. IEEE transactions on image
processing 6(5):642-652, 1997.
Taxt T, Lundervold A. Multispectral analysis of the brain
using magnetic resonance imaging. IEEE Transaction on
medical imaging 13(3):470-481, 1994.
Thune M, Olstad B, Thune N. Edge detection in noisy data
using finite mixture distribution analysis. Pattern recognition
30(5):686-699, 1997.
Wang D. A multiscale gradient algorithm for image segmentation
using watersheds, Pattern recognition 30(12):2043-2052,
1997.
Yegnanarayana M, Khemaini D. A cluster algorithm using
an evolutionary programmin-based approach. Pattern recognition
letters 18:975-986, 1997.
Yezzi A, Kichenassamy S, Kumar A, Olver P, Tannenbaum A.
A geometrical snake model for segmentation of medical imagery.
IEEE transactions on medical imaging 16(2):199-209,
1997.
Yong T, Fu K. Handbook of pattern recognition and image
processing. Academic Press, New York, 1986.

 Home
Current Issue
Table of Contents
Home
Current Issue
Table of Contents








