|
Accurate Reconstruction of
Vessels from MR Images
Juan R. Cebral(a),
Rainald Löhner(a), Peter J.Yim(b)
(a)School of Computational Sciences,
George Mason University, Fairfax,Virginia, USA
(b)Diagnostic Radiology Department,National Institutes
of Health, Bethesda, Maryland, USA
Correspondence: JR Cebral,
School of Computatinoal Sciences, George Mason University,
4400 University Drive, MS 4C7, Fairfax, VA 22030 USA. E-mail:
jcebral@gmu.edu, phone 703-993-4078, fax 703-993-4064
Abstract. Finite element calculations of arterial
hemodynamics are valuable for understanding vascular diseases,
enhancing diagnosis and surgical planning. Accurate reconstruction
of vessel anatomies is critical for computational fluid dynamics
modeling of blood flows. A methodology for constructing finite
element grids from three-dimensional medical images is presented.
The anatomical images are segmented using either a region
growing approach or deformable models. Watertight models of
bifurcating arteries are constructed by merging branches reconstructed
independently, using an adaptive voxelization scheme. The
surface model is then cut, smoothed, and optimized. Finite
element grids of tetrahedral elements are generated directly
from surface triangulations using an advancing front method.
Adaptive background grids are used to specify an element size
distribution that depends on the local curvature of the surface
model. The methodology is illustrated with several examples
including normal and diseased carotid arteries, renal arteries,
and the circle of Willis.
Keywords: Hemodynamics; Grid Generation; Magnetic Resonance
Angiography; Carotid Artery; Circle of Willis
1. Introduction
Arterial flow quantification and visualization are valuable
for determining relationships between hemodynamic factors
and progression of vascular disease [Zarins et al., 1983;
Ku et al., 1985; Berger, 1993], enhancing image-based diagnosis
[Milner et al., 1998; Steinman et al. 2000; Yim et al.,
2000], planning surgical and interventional procedures [Taylor
et al., 1999; Cebral et al., 2001], and training. Arterial
models based on idealized geometries may fail to capture
important flow features due to the strong dependence of
the flow characteristics on the vessel geometry [Moore et
al., 1999]. On the other hand, realistic patient-specific
models can provide new insights into the arterial hemodynamics
[Perktold et al., 1998; Quarteroni et al., 1998; Zhao et
al., 2000], and are required for the purposes of diagnosis
and surgical planning due to the large anatomical and physiologic
variability among individuals [Taylor et al., 1998; Cebral
et al. 2000]. Accurate reconstruction of the vessel lumen
is critical for computational fluid dynamics (CFD) modeling
of blood flow.
Vessel reconstruction from medical images is a challenging
problem due to image artifacts, noise and limited resolution.
A variety of vessel surface reconstruction algorithms have
been devised. The simplest method is iso-surface
reconstruction, in which all the vertices on the surface
share the same interpolated intensity value [Cline et al.,
1991]. However, the iso-intensity value has to be chosen
by trial-and-error and a single value may not be valid for
the entire vessel or vascular tree.
The fuzzy connectivity approach extends the conventional
region-growing method [Cebral and Löhner, 1999; Cebral and
Löhner, 2001] by assigning fuzzy membership to all points
in the image rather than the binary Inside-Outside classification
of conventional region growing. Tissue classes are identified
by placing seed points at various points inside and outside
of the vessels. The method has been found to be useful in
the visualization of MRA using intra-vascular contrast where
artery-vein overlap is considerable [Lei et al., 1999].
The marker controlled watershed segmentation method
has been applied to a high-resolution CT image of the pulmonary
arteries and a contrast-enhanced MRA of the thoracic aorta
[Yim et al., 1998; Yim and Summers, 1999]. Boundaries of
the marker-controlled watershed fall along the ridges in
the intensity surface of the gradient magnitude image. The
precise location of the watershed boundary is largely independent
of the placement of the markers that indicate the interior
and exterior of the vessel. Furthermore, smooth surfaces
can be reconstructed from the watershed segmentation without
resort to surface smoothing operations. However, image intensity
within the carotid artery in Gd contrast-enhanced MRA may
be highly inhomogeneous due to wide variation in flow rate.
Due to this problem, the watershed segmentation is prone
to large errors in vessels such as the carotid artery.
An invasive approach that combines biplane angiography
and endovascular ultrasound data has recently been used
to accurately reconstruct coronary vessels [Ilegbusi et
al., 1999; Wentzel et al., 2000]. The ultrasound images
provide cross-sectional information while the angiographic
data is used reconstruct the vessel axis. Although this
technique is able to reconstruct very accurately the lumen
of a given artery, it cannot deal properly with bifurcations
and severe stenoses [Wentzel et al., 2001].
Another methodology [Taylor et al., 2001] that has been
successfully applied to the reconstruction of the abdominal
aorta starts by probing the volumetric data with 2D slices
normal to the vessel axis. Segmentation of the vessel lumen
for each of these slices is the performed using thresholding
or level set methods. The result is a set of contours along
each arterial branch that are interpolated using analytic
curves and stacked into analytical surface patches. The
final model is the obtained by joining all the branches
using solid modeling procedures. These methods tend to localize
the largest errors at the arterial bifurcations where sharp
intersections between branches can be obtained.
Promising results have been obtained from deformable
models [Aylward et al., 1996; Frangi et al., 1999].
The surface reconstruction begins by determination of the
axis of the vessels by ridge tracking. The surface is found
by determining the radii associated with each point along
the vessel axis using estimates of scale. Radius-based surfaces
can be used as initializations for a mechanical-analog deformable
model. This method, however, is highly dependent on the
initialization of the vessel radii that may prove to be
a problem with further testing.
Recently, a new approach that combines a semi-automated
deformable model for vessel surface reconstruction and a
method for joining surface triangulations, which is necessary
at vessel bifurcations, has been presented [Yim et al.,
2001]. While this method is not completely automatic, it
avoids the problem of vessel overlapping encountered in
the region-growing schemes. The merging of surface triangulations
is made using an iso-surfacing technique combined with adaptive
background grids [Cebral et al., 2001].
In the remainder of this paper we describe the application
of region growing and deformable model segmentation for
constructing realistic arterial models from MRA images that
are used to generate finite element grids for CFD calculations
of hemodynamics.
2. Material and Methods
Given their high degree of flexibility and automation,
we have adopted unstructured grids based on tetrahedral
elements for our CFD calculations. The process of constructing
a finite element grid from medical images can be subdivided
into the following stages: model reconstruction (image processing),
solid modeling (surface processing), and grid generation.
In what follows we review each of these steps.
2.1. Region Growing Segmentation
The region growing scheme marks all the voxels topologically
connected to a given seed voxel interactively selected inside
the desired vessel, and with intensity in the same range
as the seed. The successful application of this approach
depends on the specification of an appropriate seed and
intensity range. The specification of these parameters must
be done on a trial-end-error basis for each case. However,
we have found useful to select the seed in the region with
lowest intensity within the artery, and to use the intensity
level of the seed as the lower bound of the intensity range.
If no upper bound is specified, then all the voxels connected
to the seed with intensities greater than that of the seed
are marked. The main drawback of this approach is that it
can produce leaks or false intersections between vessels
or other bright anatomical structures that are close in
space. Combining the region growing operator with image-crop
and contrast-enhancement (sharpening) operators can help
in reducing this problem. The image-crop operation
restricts all other image processing operations to a user-defined
region of interest, i.e. a bounding box in 3D. The sharpening
operator enhances the image contrast and reduces the noise
present in the image.
Once the voxels in the interior of the desired vessels
have been marked, a surface triangulation is created using
either an iso-surface extraction algorithm (marching
tetrahedra) or by direct tessellation of the
boundary surface. For further details see [Cebral and Löhner,
2001].
2.2. Segmentation with Tubular Deformable Models
The tubular deformable model procedure [Yim et al.,
2001] is based on a surface mesh that deforms towards points
of high image-gradient intensity while maintaining smoothness
of the surface. It is based on a coordinate system that
is tubular in nature and that allows for curvature of the
cylindrical axis. In contrast to previous models [Frangi
et al., 1999], the vertices deform only in the radial direction
and their position is described only by their radial location.
This formulation of the deformation process eliminates bias
towards forming tubes of smaller or larger diameter.
The deformation process is straightforward. First the location
of each vertex is initialized to be the point along its
radial path where the maximum of the gradient magnitude
occurs. The gradient image is obtained by convolution of
the image with the gradient of the normalized spherical
Gaussian function. The radial function then deforms so as
to minimize discontinuities in the vertex radial position
along the surface while maximizing the proximity of the
vertices to edges in the image. The deformations are applied
simultaneously to all vertices and repeated until an equilibrium
condition is obtained.
The cylindrical deformation process could be applied directly
to the modified cylindrical coordinate system. However,
bunching of vertices will occur where radial lines merge.
Where such bunching occurs, the effective elasticity of
the surface becomes significantly greater which reduces
the surface smoothness. This effect is removed by merging
vertices in the deformation process to match the merging
of the radial lines. This precludes two vertices from co-existing
on the same radial line.
In practice, the axis of the tubular deformable model is
defined from a sequence of points along the center of the
vessel identified by the user. Alternatively, a skeletonization
schemes can be used to automate the process of vessel axis
definition [Yim et al., 2000; Choyke et al., 2001]. For
bifurcations, the surface is reconstructed by two applications
of the procedure: one for the parent vessel and its continuation
onto one branch and the second for the parent vessel and
its continuation onto the second branch. We have found this
method to be highly reliable in qualitative and quantitative
studies.
2.3. Solid Modeling
The generation of finite element grids requires a watertight
definition of the computational domain, i.e. a surface model
with no holes, gaps, self-intersections or overlapping components.
Traditionally, analytical surface patches such as Coons
or NURBS have been fitted to the reconstructed triangulation.
Boolean operations between different components, e.g. to
merge different arterial branches, can then be performed
using available geometric modeling packages. Alternatively,
we generate finite element grids directly from the surface
triangulations. This implies further processing of the reconstructed
models.
Boolean operations between surface triangulations are carried
out using a volumetric technique [Cebral et al., 2001].
The basis of the method is the extraction of the zero-level
iso-surface of a (signed) distance map defined on an adaptive
background grid that covers the entire computational domain.
The distance map is computed as the shortest distance from
a background grid point to the domain surface. This methodology
has been applied to merge carotid artery branches
reconstructed independently using the tubular deformable
model.
Small-scale imperfections in the reconstructed triangulation
are then filtered using a non-shrinking surface smoothing
algorithm [Taubin, 1995]. The basis of this scheme is the
iterative application of a Laplacian operator, which is
equivalent to the calculating the mechanical equilibrium
of spring system between neighboring surface nodes. A modified
version of this algorithm that considers forces between
not only nearest neighbors of a given point but also other
close points has been used to avoid self-intersections in
vessels of small diameters [Cebral et al., 2000].
Several surface cutting operations can be used to
cut arterial branches at selected locations. These cuts
are performed in order to restrict the CFD analysis to a
desired portion of the vessel tree. A method for cutting
vessel branches along contours of geodesic distances to
a given source point yields good results in regions far
from bifurcations [Cebral et al., 2000]. These cuts tend
to be normal to the vessel axis and can be restricted to
one branch at a time (topological constraint). The point
where the cut is performed can be specified interactively
with only one mouse click. When this method fails to provide
the desired results, planar cuts restricted by a bounding
box are used. The cutting plane and the bounding box are
both graphically defined by the user.
Before proceeding to the finite element grid generation,
the quality of the surface triangulation is improved using
a mesh optimization algorithm. Duplicated points,
i.e. points that are very close in space, and degenerated
elements, i.e. elements with a repeated node, are first
identified and deleted. Large aspect ratio or very small
elements are then removed from the triangulation using an
edge-collapsing algorithm. Diagonal swaps are then performed
between adjacent triangles in order to minimize the maximum
angle of the triangular elements. These operations are prevented
if the new configuration has a significantly different surface
normal than the original configuration. For further details
see [Cebral and Löhner, 2001].
2.4. Grid Generation
As mentioned above, we have developed algorithms to generate
tetrahedral finite element grids directly from the reconstructed
models [Löhner, 1996]. These procedures do not need an analytical
surface representation of the computational domain. Rather,
they operate directly on the discrete data, i.e.
surface triangulation. We have found that this approach
yields accurate surface grids and can be fully automated.
Given a desired element size distribution, the generation
of a volumetric grid composed of tetrahedral elements begins
by first creating a new triangulation of the domain surface.
We have developed an advancing front method [Löhner, 1996;
Löhner, 1997; Owen, 1998] that creates a surface mesh using
the reconstructed model as a support surface. A feature
detection algorithm is used to identify and preserve
surface features such as ridges (i.e. edges shared by triangles
with large deviation of their unit normals), and corner
points (i.e. points connected to one or more than two ridges).
Linear or quadratic interpolation is used to place
the newly created points on the support surface. Topological
constraints are used during the interpolation in order
to avoid jumps between locally unconnected surface elements,
e.g. between arterial branches that are very close in space.
The specification of the element size distribution is typically
done using background grid and sources. In many cases, such
as carotid arteries of normal subjects, using a uniform
element distribution is usually acceptable. In these cases,
the element size is selected in order to have a minimum
number of elements across the smallest cross section of
the model. However, in cases with geometric features of
different scales, such as arteries with high degree of stenosis
or larger vessel trees, this approach may result in a very
large number of elements and therefore extremely long computing
times. For this reason we have developed methods to specify
element sizes based on the local curvature of the surface
model, using adaptive background grids [Löhner and
Baum, 1992]. The basic idea of the methodology is to refine
the background grid close to the model surface and specify
element sizes at the nodes of this mesh based from the local
curvature of the closest surface. This approach tends to
yield grids that contain roughly the same number of elements
across any cross-section of the model.
3. Results
In this section we present several examples that illustrate
the process of generating finite element grids for computational
fluid dynamics calculations of hemodynamics from medical
images.
3.1. Carotid Arteries
The methodology was first applied to the construction of
realistic models of the carotid bifurcation of human subjects
from Gd contrast-enhanced MRA images. The first of these
cases corresponds to a normal subject. Fig. 1a shows the
maximum intensity projection (MIP) of the anatomical images.
The reconstructed model of the right carotid artery is shown
in Fig. 1b. Deformable models were used to reconstruct the
internal carotid artery (ICA) and the external carotid artery
(ECA) independently. These branches were then merged using
the adaptive voxelization method in order to create a watertight
model of the carotid bifurcation. A detail of the reconstructed
models of the ICA and ECA extending into the common carotid
artery (CCA) are shown in Fig. 1c. The merged model of the
bifurcation is shown in Fig. 1d. This model was then used
as a support surface to generate a finite element grid of
tetrahedral elements with a uniform element size distribution.
The surface of the finite element mesh is shown in Fig.
1e.
|
|
 |
a)
c) d) e)
Figure 1. Realistic
carotid bifurcation model from MRA images of a normal subject:
a) MIP projection of the MRA images; b) reconstructed model
of the right carotid artery; c) model of the ICA and ECA
extending into the CCA; d) bifurcation model after merging;
e) surface of the final finite element grid.
The second case corresponds to the reconstruction of the carotid
bifurcation from contrast-enhanced MRA images of a patient
with atherosclerosis. As in the previous case, deformable
models were used to reconstruct each arterial branch independently
in order to avoid intersections with other close vessels present
in the images. These branches were then merged into a watertight
model and used as a support surface to generate a finite element
grid with a uniform element size distribution. The MIP of
the anatomical images is shown in Fig. 2a, the reconstructed
model of the right carotid artery is shown in Fig. 2b and
the surface of the final finite element grid is shown in Fig.
2c.
Figure 2. Construction
of realistic carotid bifurcation model from MRA images of
a patient with atherosclerosis: a) MIP projection of the
contrast-enhanced MRA images; b) reconstructed model of
the right carotid artery; c) surface of the final finite
element grid.
A detail of the reconstructed models of the ICA and ECA
extending into the CCA are shown in Fig. 3a and 3b, respectively.
The surface triangulation of the bifurcation model after
merging the ICA and ECA models is shown in Fig. 3c.
3.2. Renal Arteries
Tubular deformable models were used to reconstruct a portion
of the abdominal aorta in the region of the renal arteries
from contrast-enhanced MRA images (Fig. 4a). A deformable
model was used for each branch and later all the branches
were merged using the adaptive voxelization method. The
watertight model thus created (Fig. 4b) was then used to
generate a finite element grid with a uniform element size
distribution (Fig. 4c).
a)
b) c)
Figure 3. Merging
models of the ICA and ECA extending into the CCA in order
to create a watertight model of the carotid bifurcation:
a) model of the ICA; b) model of the ECA; c) merged bifurcation
model.
a)
b)
c)
Figure 4. Generation
of a finite element grid of the renal arteries from contrast-enhanced
MRA images: a) MIP of the anatomical images; b) reconstructed
model; c) finite element mesh.
3.3. Circle of Willis
The region growing segmentation approach was applied to
the construction of realistic anatomical models of the circle
of Willis from 3D MRA images. The MIPs of the anatomical
images before and after applying the region growing procedure
are shown in Fig. 5a and 5b, respectively. The reconstructed
model after smoothing and optimization is shown in Fig.
5c.
a)
b) c)
Figure 5. Reconstruction
of the circle of Willis from MRA images: a) MIP of original
images; b) MIP after region growing; c) reconstructed surface
model.
a) b)
Figure 6. Finite element
grid of the circle of Willis: a) final geometrical model
after cutting arterial branches;
b) surface of the final finite element grid with uniform
element size distribution.
The final geometrical model after cutting several arterial
branches along contours of geodesic distances is shown in
Fig. 6a. This model was then used to generate a finite element
grid with uniform element size distribution. The final mesh
(Fig. 6b) contained approximately 4 million elements.
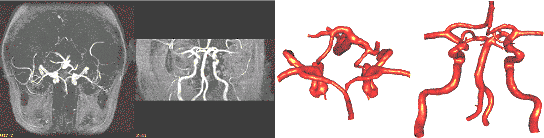
A second case illustrates the use of adaptive background
grids to specify an element size distribution that depends
on the local surface curvature. Fig. 7a and 7b show the
MIP of the MRA images from two different viewpoints. The
reconstructed model is shown from similar viewpoints in
Fig. 7c and 7d.
a) b)
c) d)
Figure 7. Circle of
Willis model reconstructed from MRA images: a) and b) MIP
projections from different viewpoints; c) and d) reconstructed
model from similar viewpoints.
A close-up view of the triangulation of the final model
is shown in Fig. 8a. This triangulation was then used to
refine an adaptive background grid based on its local curvature
in order to specify an element size distribution that yields
roughly the same number of elements across any cross-section
of the model. The generated finite element grid (Fig. 8b)
contained approximately 1 million elements.
a)
b)
Figure 8. Finite element
mesh of the circle of Willis: a) surface triangulation of
the final geometrical model; b) surface of the finite element
grid generated using adaptive background grids to specify
an element size distribution that depends on the local surface
curvature.
4. Discussion
A methodology for generating finite element grids from
medical images has been presented. The images are first
segmented using either a region growing approach followed
by an iso-surface extraction or using tubular deformable
models. In the case of deformable models, each arterial
branch is reconstructed independently, avoiding problems
of intersections with other close anatomical structures,
and then merged into a watertight surface using an adaptive
voxelization scheme. The reconstructed triangulation is
then cut at desired locations, smoothed and optimized in
order to improve the element quality. This final model is
the used as a support surface for unstructured grid generation.
A feature detection algorithm is combined with an advancing
front method to generate a new surface triangulation with
the desired element size distribution. This triangulation
is then used as the initial front to generate a volumetric
finite element grid of tetrahedral elements using an advancing
front technique. Adaptive background grids are used in order
to specify curvature-dependent element size distributions
that result in volumetric grids with a roughly uniform number
of elements in any cross-section of the model.
The region growing approach requires the proper selection
of a seed voxel and an intensity range. This has to be done
on a trial-and-error basis. However, quite complex vascular
trees can be reconstructed using this approach. Another
limitation of the method is that for a given intensity threshold
intersections between close vessels can be obtained. One
of such intersections was generated in the model of the
circle of Willis shown in Fig. 5c. This false intersection
was then discarded when the model was cut at several locations.
The tubular deformable model is a very reliable semi-automatic
method for reconstructing arterial branches independently.
Due to its intrinsic smoothing constraints, false intersections
with other close vessels are not generated. This approach
requires the specification of the arterial axis, which can
be done manually or using skeletonization algorithms. Another
limitation of this method is that pathological entities
that are not tubular in nature, such as aneurysms, will
not be properly treated.
The grid generation from surface triangulations has the
advantage of avoiding the need of splitting the surface
into patches and fitting analytical functions. However,
this approach requires several surface processing operations
such as merging, smoothing, cutting and optimization. These
operators also allow for the possibility of using different
segmentation techniques for different entities, such as
vessels and aneurysms, which can then be combined into a
single model.
The use of adaptive background grids to specify curvature-dependent
element size distributions is very useful to reduce the
total number of elements in the final grids, while maintaining
a reasonable mesh resolution everywhere. Using uniform element
size distributions in cases with large variations in spatial
scales would result in several millions of elements, and
therefore very long computing times.
The methodology has been previously validated using a glass
phantom of the carotid artery with stenosis [Cebral at al.,
2001]. Errors in the reconstructed radii were less than
one percent. We intend to further evaluate this methodology
using in vivo data from different imaging modalities.
References
Aylward S, Bullit E, Pizer S, Eberly D. Intensity ridge
and widths for tubular object segmentation and description.
In proceedings of the IEEE Workshop, Mathematical Methods
in Biomedical Image Analysis, June 1996.
Berger, SA. Flow in large vessels, in Contemporary Mathematics:
Fluid Dynamics in Biology, Cheer AY, van Dam CP, Editors.
American Mathematical Society, 1993, 141: 479-518.
Cebral JR, Löhner R. From medical images to CFD meshes.
In proceedings of the 8th International Meshing
Roundtable, South Lake Tahoe, California, October 10-13,
1999, 321-332.
Cebral JR, Löhner R, Burgess JE. Computer simulation of
cerebral artery clipping: relevance to aneurysm neuro-surgery
planning. In proceedings of ECCOMAS 2000, John Wiley and
Sons, Barcelona, Spain, September 11-14, 2000.
Cebral JR, Löhner R, Choyke PL, Yim PJ. Merging of intersecting
triangulations for finite element modeling. Journal of
Biomechanics, in press 2001.
Cebral JR, Löhner R. From medical images to anatomically
accurate finite element grids. International Journal
for Numerical Methods in Engineering, 51:985-1008, 2001.
Cebral JR, Yim PJ, Löhner R, Soto O, Marcos H, Choyke PL.
New methods for computational fluid dynamics modeling of
carotid artery from magnetic resonance angiography. In proceedings
of SPIE Medical Imaging 2001: Physiology and Function from
Multidimensional Images, 2001, 4321: 177-187.
Cebral JR, Löhner R, Soto O, Choyke PL, Yim PJ. Patient-specific
simulation of carotid artery stenting using computational
fluid dynamics. In proceedings of MICCAI 2001, Utretch,
Netherlands, October 2001.
Choyke PL, Yim PJ, Marcos H, Mullick R, Summers R, Ho V.
Hepatic MR angiography: a multiobserver comparison of visualization
methods. American Journal of Roentgenology, 176:465-470,
2001.
Cline HE, Lorensen WE, Souza SP, Jolesz FA, Kikinis R,
Gerig G, Kennedy TE. 3D surface rendered MR images of the
brain and its vasculature. Journal of Computer Assisted
Tomography, 15:344-351, 1991.
Frangi AF, Niessen WJ, Hoogeveen RM. Model-based quantitation
of 3-D magnetic resonance angiographic images. IEEE Transactions
on Medical Imaging, 18:946-956, 1999.
Ilegbusi OJ, Hu Z, Nesto R, Waxman S, Cyganski D, Kilian
J, Stone PH, Feldman CL. Determination of blood flow and
endothelial shear stress in human coronary artery in vivo.
Journal of Invasive Cardiology, 11(11):667-674, 1999.
Ku DN, Gibbens DP, Zarins CK, Glagov S. Pulsatile flow
and atherosclerosis in the human carotid bifurcation: positive
correlation between plaque location and low and oscillating
shear stress. Atherosclerosis, 5:293-302, 1985.
Lei T, Udupa JK, Saha PK, Odhner D. MR Angiographic visualization
and artery-vein separation. In proceedings of SPIE Medical
Imaging, 1999, 3658:58-66.
Löhner R, Baum JD. Adaptive h-refinement on 3-d unstructured
grids for transient problems. International Journal for
Numerical Methods in Fluids, 14:1407-1419, 1992.
Löhner R. Regridding surface triangulations. Journal
of Computational Physics, 126:1-10, 1996.
Löhner R. Progress in grid generation via the advancing
front technique. Engineering with Computers, 12:86-210,
1996.
Löhner R. Automatic unstructured grid generators. Finite
Elements in Analysis and Design, 25:111-134, 1997.
Milner JS, Moore JA, Rutt BA, Steinman DA. Hemodynamics
of human artery bifurcations: computational studies with
models reconstructed from magnetic resonance imaging of
normal subjects. Journal of Vascular Surgery, 27:143-156,
1998.
Moore JA, Steinman DA, Holdsworth DW, Ethier CR. Accuracy
of computational hemodynamics in complex arterial geometries
reconstructed from magnetic resonance imaging. Annals
of Biomedical Engineering, 27:32-41, 1999.
Owen SJ. A survey of unstructured mesh generation technology.
In proceedings of the 7th International Meshing Roundtable,
Sandia National Lab., 1998, 239-267.
Perktold K, Hofer M, Karner G, Trubel W, Schima H. Computer
simulation of vascular fluid dynamics and mass transport:
optimal design of arterial bypass anastomoses. In proceedings
of ECCOMAS 98, John Wiley & Sons, 1998, 2:484-489.
Quarteroni A, Tuveri M, Veneziani A. Computational vascular
fluid dynamics: problems, models and methods. Computing
and Visualization in Science, 2:163-197, 1998.
Steinman DA, Poepping TL, Tambasco M, Rankin RN, Holdsworth
DW. Flow patterns at the stenosed carotid bifurcation: effect
of concentric versus eccentric stenosis. Journal of Biomechanics,
28:415-423, 2000.
Taubin G. A signal processing approach to fair surface
design. In proceedings of Computer Graphics, 1995, 351-358.
Taylor CA, Hughes TJR, Zarins CK. Finite element modeling
of blood flow in arteries. Computer Methods in Applied
Mechanics and Engineering, 158:155-196, 1998.
Taylor CA, Draney MT, Ku JP, Parker D, Steele BN, Wang
K, Zarins CK. Predictive medicine: computational techniques
in therapeutic decision-making. Computer Assisted Surgery,
4:231-247, 1999.
Taylor CA, Parker D, Wang KC. Image based geometric modeling
of the human aorta. In proceedings of the ASME-BED Bioengineering
Conference, vol. 50, 2001, 623-624.
Wentzel JJ, Whelan DM, van Der Giessen WJ, van Beusekom
HMM, Andhyiswara I, Serruys PW, Slager CJ and Kram R. Coronary
stent implantation changes 3d vessel geometry and 3d shear
stress distribution. Journal of Biomechanics, 33:1287-1295,
2000.
Wentzel JJ, Krams R, Schuurbiers JC, Oomen JA, Kloet J,
van Der Giessen WJ, Serruys PW, Slager CJ. Relationship
between neointimal thickness and shear stress after wallstent
implantation in human coronary arteries. Circulation,
103(13):1740-5, 2001.
Yim PJ, Kim D, Lucas C. High resolution four-dimensional
surface reconstruction of the right heart and pulmonary
arteries. In proceedings of SPIE Medical Imaging, 1998,
3338:726-738.
Yim PJ, Summers MR. Analytic surface reconstruction by
local threshold estimation in the case of simple intensity
contrasts. In proceedings of SPIE Medical Imaging,
1999, 3660:288-300.
Yim PJ, Mullick R, Summers RM, Marcos H, Cebral JR, Löhner
R, Choyke PL. Measurement of stenosis from magnetic resonance
angiography using vessel skeletons. In proceedings of
SPIE Medical Imaging, 2000.
Yim PJ, Summers RM, Choyke PL. Gray-scale skeletonization
of small vessels in magnetic resonance angiograms. IEEE
Transactions on Medical Imaging, 19:568-576, 2000.
Yim PJ, Cebral JR, Mullick R, Choyke PL. Vessel surface
reconstruction with a tubular deformable model. Submitted
to IEEE Transactions on Medical Imaging, 2001.
Zarins CK, Giddens DP, Bharadvaj BK, Sottiurai VS, Mabon
RF, Glagov S. Carotid bifurcation atherosclerosis. quantitative
correlation of plaque localization with flow velocity profiles
and wall shear stress. Circulation Research, 53(4):502-514,
1983.
Zhao SZ, Xu XY, Hughes AD, Thom SA, Stanton AV, Ariff B,
Long Q. Blood flow and vessel mechanics in a physiologically
realistic model of a human carotid arterial bifurcation.
Journal of Biomechanics, 33: 975-984, 2000.
| 
 Home
Current Issue
Table of Contents
Home
Current Issue
Table of Contents